Hi all,
I’m Zak, a PhD student at the Peter Doherty Institute for Infection and Immunity, Melbourne, Australia. My PhD project is focused on using gene editing tools, CRISPR/Cas9 base editors (BEs) and CRISPR/Cas13b to target Hepatitis B virus (HBV) DNA and RNA respectively to reduce protein expression and replication. The long-term aim of this project is to develop a potential novel therapeutic using these gene editing tools.
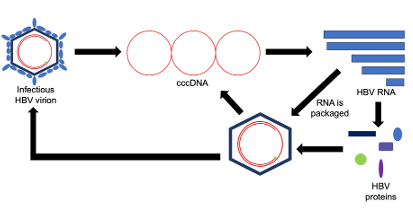
The HBV replication cycle is composed of both DNA and RNA intermediates. HBV is mainly a DNA virus, with the main DNA form being covalently closed circular DNA (cccDNA). This cccDNA is then read by the host cell to produce multiple different RNAs. These RNAs are subsequently read by the host cell to produce HBV proteins. One of these RNAs is also packaged by HBV proteins to form infectious HBV viral particles or form more cccDNA.
Figure 1: HBV replication cycle.
Both HBV DNA and RNA are not targeted by any current therapy, making them an attractive target for a potential new therapy to improve cure rates.
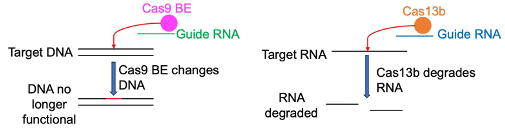
In our group, I mainly focus on targeting HBV DNA using the gene editing tool, CRISPR/Cas9 BEs. Traditional CRISPR/Cas9 is utilised by bacteria to target invading pathogens’ DNA and prevent infection of the bacteria. CRISPR/Cas9 is comprised of a Cas9 protein and a guide RNA which contains a variable sequence. This variable sequence matches with the target DNA. The guide RNA binds to the Cas9 protein and guides it to the target site. Following recognition of the target sequence and other essential motifs, the Cas9 breaks both DNA strands and can make the DNA non-functional. We can change the variable sequence so that Cas9 can target HBV DNA. A new form of CRISPR/Cas9 has been developed, these are called BEs. BEs utilise the Cas9 guiding system but instead of breaking both DNA strands, BEs introduce specific DNA base changes into target DNA. This approach is more specific and therefore potentially safer than traditional Cas9. We can use this BE to introduce a mutation that can lead to a truncated and subsequently non-functional HBV protein.
To target HBV RNA, we can use another gene editing tool, CRISPR/Cas13b. Cas13 works in a similar manner as traditional CRISPR/Cas9 but instead targets RNA for degradation. Our lab was the first lab in the world to utilise Cas13 to target HBV RNA.
Figure 2: Mechanism of CRISPR/Cas9 BE and CRISPR/Cas13b
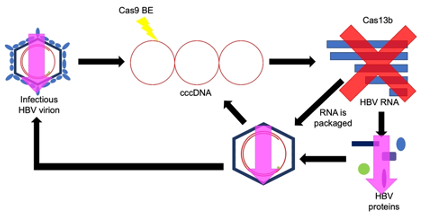
During my PhD, I have designed Cas9 BEs guide RNAs to target different regions of the HBV genome that would create truncated HBV proteins. I tested which of these guides was best at reducing HBV replication and protein expression in cell culture models in the laboratory. After finding the best Cas9 BE guide RNA, I then co-delivered this guide with Cas9 BE, a previously determined best Cas13b guide and Cas13b together and measured HBV replication and protein expression in cell culture models. A combination of Cas9 and Cas13 has never been tested before. Preliminary results showed that Cas9 BE and Cas13b can work together to reduce protein expression and replication, with minimal interference between the two gene editing tools. We will next perform HBV RNA analysis and mice experiments.
Figure 3: Cas9 BE and Cas13b targeting HBV DNA and RNA respectively reduced HBV
proteins and DNA.
These studies will determine the utility of Cas9 BEs in combination with Cas13, to simultaneously target HBV DNA and RNA aspects of the viral ‘life cycle’, a combination never tested before.